|
Like bacteria, archaea are prokaryotes (micro-organisms with neither a nucleus with a distinct membrane nor specialized organelles). Although they resemble bacteria morphologically, archaea have genes and metabolic pathways similar to those of eukaryotes (organisms whose cells have a distinct nucleus) and their cell membranes have high levels of glycerol-ether lipids, whereas cell membranes of bacteria and eukaryotes comprise mostly glycerol-ester lipids (Dridi et al., 2011). Because of their glycerol-ether lipids, archaeal cells are more difficult to break open than those of bacteria and, consequently, are likely to be under-represented in many DNA-based microbiota/microbiome studies (Dridi et al., 2009). There are five phyla of archaea (Euryarchaeota, Crenarchaeota, Nanoarchaeota, Xenarchaeota/Korachaeota, Thaumarchaeota), but only representatives of the Euryarchaeota are frequently found in the human gut microbiota, with methanogenic archaea predominating in this environment. The archaeal component of the gut microbiota is sometimes referred to as the archaeome. Until recently it was thought all archaea of the human gut microbiota were obligate anaerobes, but it is now possible to isolate some archaea aerobically from gut samples if they are grown with an external hydrogen (H2) source or in high salt conditions (Lagier et al., 2016). In addition to the methanogenic and halophilic archaea mentioned below, molecular studies have suggested the presence of members of the orders Methanosarcinales, Thermoplasmatales, Methanomicrobiales and Nitrososphaerales in the human gut microbiota, but none of these micro-organisms has been isolated (Dridi et al., 2011; Gaci et al., 2014). In many instances isolation and cultivation of archaea is complicated by their fastidious atmospheric requirements [e.g. 20 % CO2, 80 % H2, 1–25 bar (14.5–36 psi) for some of the methanogenic archaea] (Dridi et al., 2011). Methanogens Methanogens are archaea that generate methane (CH4) from a limited number of simple substrates [e.g. carbon dioxide (CO2); H2; formate (CHOO–); trimethylamine (TMA; N(CH3)3); methanol (CH3OH)] produced when dietary substrates and other organic matter are broken down by hydrolytic and fermentative microbes in the gut (Gaci et al., 2014; Figure 1). These simple substrates are considered the ‘end products’ of microbial catabolism. Between 30 and 40 % of H2 and CH4 produced in the intestine is absorbed by the colonic mucosa and taken up into the bloodstream, with the rest expelled from the body through belching or flatulence (Gibson et al., 1993; Carbonero et al., 2012; Figure 1). It has been suggested humans acquire methanogenic archaea via environmental contamination; once these microbes find favourable physicochemical and nutrient conditions in the gut, they colonize the intestine and become established members of the gut microbiota (Dridi et al., 2011). Breath tests suggest between a third and two-thirds of healthy adults produce CH4 (Levitt et al., 2006), while sensitive molecular-based studies show most adults carry methanogenic archaea in their intestine (>80–96 %: Mihajlovski et al., 2010; Dridi et al., 2009; Dridi et al., 2012a; Vanderhaeghen et al., 2015). Abundance of methanogenic archaea increases from the ascending (proximal) to the descending (distal) colon (0.003 % to 11 % of total gut prokaryotes; Dridi et al., 2011), and increases from childhood into adulthood and old age (Mihajlovski et al., 2010). The hydrogenotrophic (H2-utilizing) archaea belonging to the order Methanobacteriales (Methanibrevibacter, Methanosphaera) are the most frequently detected (89 % and 65 % of adults and children, respectively) and most abundant methanogens in human faeces (0.52 % and 0.15 % of the total faecal microbiota of adults and children, respectively) (Vanderhaeghen et al., 2015). Hydrogenotrophic methanogens use H2 for the reduction of CO2 and methyl-compounds. By removing H2 from the gastrointestinal tract, these archaea optimize fermentation in this environment and modify the metabolic pathways of fermentative bacteria (Gaci et al., 2014). Reduction of CO2 using H2 efficiently reduces the gas partial pressure in the colon (4 H2 + CO2 → CH4 + 2 H2O). The methylotrophic (one-carbon-compound-utilizing) members of the order Methanomassiliicoccales (Methanomassiliicoccus) are present in 50 % of adults, with prevalence seemingly increasing from infancy to adulthood (Dridi et al., 2012a; Vanderhaeghen et al., 2015). 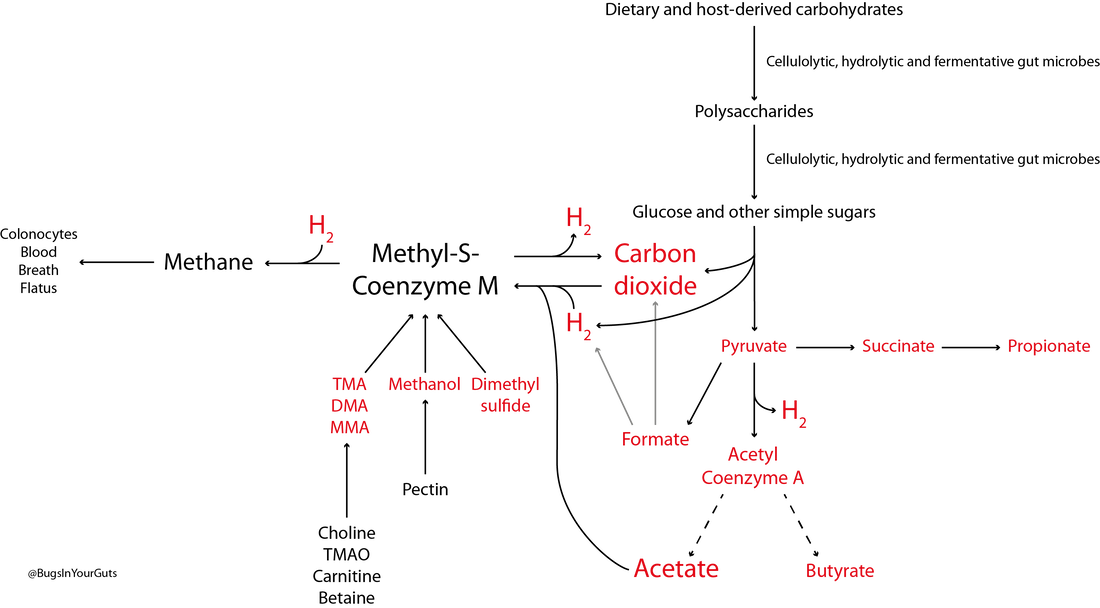 Figure 1. Microbial fermentation and hydrolysis of dietary substrates [predominantly polysaccharides (10–60 g/day) and proteins (6–18 g/day)] in the gastrointestinal tract leads to the release of end-products. Methanogens, therefore, have a terminal position in microbial trophic chains. The fate of carbohydrates, methylamines, methanol and dimethyl sulfide are shown in the figure. Broken arrows, incomplete pathways shown; full details can be found in the references cited below. In addition to those shown in red, ethanol, lactate, amino acids, ammonia, sulfate, phenolic compounds, indoles and branched-chain fatty acids are end-products of microbial catabolism released into the intestinal lumen. Methanobrevibacter smithii uses carbon dioxide (CO2) (or formate) and hydrogen (H2) to produce methane. Methanosphaera stadtmanae, members of the order Methanomassiliicoccales and possibly Methanobrevibacter smithii use methanol and H2 to produce methane. Members of the order Methanomassiliicoccales use methylamines (TMA, trimethylamine; DMA, dimethylamine; MMA, monomethylamine) and H2 to produce methane. Acetate to methane, aceticlastic methanogenesis. Methylamines, methanol and dimethyl sulfide to methane, methylotrophic methanogenesis. CO2 to methane: hydrogenotrophic methanogenesis. Information taken from Borrel et al. (2013a), Carbonero et al. (2012), Dridi et al. (2011), Gaci et al. (2014) and Chaudhary et al. (2015). Methanobrevibacter smithii Methanobrevibacter smithii is the most abundant archaeon in the human gut, and is detected in almost all adults (Mihajlovski et al., 2010). It can colonize the gut from the caecum to the rectum (Gaci et al., 2014). Cultivation work has shown it represents between 0.001 and 13 % of total anaerobes in human faeces (Miller & Wolin, 1982). Using a protocol optimized for the extraction and detection of archaeal DNA in faeces, it was shown that 95.5 % of individuals carry Methanobrevibacter smithii (Dridi et al., 2009). The archaeon has not been detected in the faeces of newborns (Mihajlovski et al., 2010), but has been detected in the faeces of children as young as 2 weeks of age (Dridi et al., 2009). It reduces CO2 to CH4 with H2 as the primary electron donor, and can use formate directly as a substrate to produce CH4 (Jones et al., 1987; Chaudhary et al., 2015; Vanderhaeghen et al., 2015). It can also use ammonium (NH4+) and ammonia (NH3) as sources of nitrogen (N) and consume ethanol (CH3CH2OH) present in the intestine, contributing to non-methanogenic removal of bacterial end-products of fermentation (Samuel et al., 2007; Hansen et al., 2011). The cell surface of Methanobrevibacter smithii is covered with carbohydrates (glycans) that mimic those found in the intestinal mucosa, facilitating its colonization of the intestine (Samuel et al., 2007). Methanobrevibacter smithii encodes a bile salt hydrolase that can degrade both tauro and glycol conjugated bile acids, thereby contributing to bile acid modification in the human gut (Jones et al., 2008). Methanomassiliicoccus luminyensis Methanomassiliicoccus luminyensis is currently the only cultivated member of the genus Methanomassiliicoccus. It was first isolated from human faeces on methanol and H2 (Dridi et al., 2012b). This methylotrophic archaeon has an energy metabolism distinct from other cultured methanogens: it can use methanol or TMA as an electron acceptor (and may be able to use dimethyl sulfide) to produce CH4 (Dridi et al., 2012b; Brugère et al., 2014). The ability of of Methanomassiliicoccus luminyensis to remove TMA from the gut has potential health benefits to humans, leading to the proposed use of archaebiotics in individuals with fish odour syndrome and those at risk of cardiovascular disease (Brugère et al., 2014). ‘Candidatus Methanomassiliicoccus intestinalis’ and ‘Candidatus Methanomethylophilus alvus’, related to Methanomassiliicoccus luminyensis, have been enriched from human faeces using methanol; both species are predicted to use methylamines to produce CH4. Although the draft genome sequences of these archaea are available, neither has been successfully isolated and preserved, hence their Candidatus status (Borrel et al., 2012; Borrel et al., 2013b). In addition to the human gut microbiota, 16S rRNA gene and/or mcrA (methyl:coenzyme M reductase, a functional marker gene of methanogens) sequences of members of the order Methanomassiliicoccales have been found in the gastrointestinal tract of the elephant, giant tortoise, tortoise, rat, cattle, sheep, yak, reindeer and wallaby, and the human oral cavity (Söllinger et al., 2016). Methanosphaera stadtmanae Methanosphaera stadtmanae is detected in up to a third of western adults, with abundance appearing to increase from adulthood into old age (Dridi et al., 2009; Mihajlovski et al., 2010). It only produces CH4 by reducing methanol (derived from microbial degradation of pectin) in the presence of H2, and needs acetate (CH3COOH) and CO2 as carbon sources (Fricke et al., 2006). Growth of Methanosphaera stadtmanae is enhanced in the presence of tungsten (Dridi et al., 2012c). In vitro, Methanosphaera stadtmanae and Methanobrevibacter smithii have been shown to form biofilms on various substrates; however, it is not know if it they contribute to mucosa-associated biofilms in the human gut (Bang et al., 2014a). In addition, both archaea have been shown to stimulate dendritic cells and peripheral blood mononuclear cells in vitro, suggesting they may be specifically recognized by the human innate immune system, though this has not been demonstrated in vivo (Bang et al., 2014b; Blais Lecours et al., 2014). Halophilic archaea As their name implies, halophiles are salt-loving organisms able to thrive in high salt concentrations (slight halophiles, 0.3–0.8 M NaCl; moderate halophiles, 0.8–3.4 M NaCl; extreme halophiles, 3.4–5.1 M NaCl). The human gut is not a salty environment (135–145 mM sodium), but moderatey halophilic, aerobic archaea belonging to the order Halobacteriales have been isolated in small numbers from intestinal mucosal samples taken from patients with inflammatory bowel disease (Oxley et al., 2010). More recently, the halophilic archaea Haloferax alexandrines and ‘Haloferax massiliensis‘ have been isolated from the human gut, but their role(s) and abundance in this environment are not yet known (Khelaifia & Raoult, 2016; Lagier et al., 2016). References Bang, C., Ehlers, C., Orell, A., Prasse, D., Spinner, M., Gorb, S. N., Albers, S. V. & Schmitz, R. A. (2014a). Biofilm formation of mucosa-associated methanoarchaeal strains. Front Microbiol 5, 353. Bang, C., Weidenbach, K., Gutsmann, T., Heine, H. & Schmitz, R. A. (2014b). The intestinal archaea Methanosphaera stadtmanae and Methanobrevibacter smithii activate human dendritic cells. PLoS One 9, e99411. Blais Lecours, P., Marsolais, D., Cormier, Y., Berberi, M., Haché, C., Bourdages, R. & Duchaine, C. (2014). Increased prevalence of Methanosphaera stadtmanae in inflammatory bowel diseases. PLoS One 9, e87734. Borrel, G., Harris, H. M., Tottey, W., Mihajlovski, A., Parisot, N., Peyretaillade, E., Peyret, P., Gribaldo, S., O’Toole, P. W. & Brugère, J. F. (2012). Genome sequence of “Candidatus Methanomethylophilus alvus” Mx1201, a methanogenic archaeon from the human gut belonging to a seventh order of methanogens. J Bacteriol 194, 6944-6945. Borrel, G., O’Toole, P. W., Harris, H. M., Peyret, P., Brugère, J. F. & Gribaldo, S. (2013a). Phylogenomic data support a seventh order of methylotrophic methanogens and provide insights into the evolution of methanogenesis. Genome Biol Evol 5, 1769-1780. Borrel, G., Harris, H. M., Parisot, N., Gaci, N., Tottey, W., Mihajlovski, A., Deane, J., Gribaldo, S., Bardot, O., Peyretaillade, E., Peyret, P., O’Toole, P. W. & Brugère, J. F. (2013b). Genome sequence of “Candidatus Methanomassiliicoccus intestinalis” Issoire-Mx1, a third Thermoplasmatales-related methanogenic archaeon from human feces. Genome Announc 1, pii:e00453-13. Brugère, J. F., Borrel, G., Gaci, N., Tottey, W., O’Toole, P. W. & Malpuech-Brugère, C. (2014). Archaebiotics: proposed therapeutic use of archaea to prevent trimethylaminuria and cardiovascular disease. Gut Microbes 5, 5-10. Carbonero, F., Benefiel, A. C. & Gaskins, H. R. (2012). Contributions of the microbial hydrogen economy to colonic homeostasis. Nat Rev Gastroenterol Hepatol 9, 504-518. Chaudhary, P. P., Gaci, N., Borrel, G., O’Toole, P. W. & Brugère, J. F. (2015). Molecular methods for studying methanogens of the human gastrointestinal tract: current status and future directions. Appl Microbiol Biotechnol 99, 5801-5815. Dridi, B., Henry, M., El Khéchine, A., Raoult, D. & Drancourt, M. (2009). High prevalence of Methanobrevibacter smithii and Methanosphaera stadtmanae detected in the human gut using an improved DNA detection protocol. PLoS One 4, e7063. Dridi, B., Raoult, D. & Drancourt, M. (2011). Archaea as emerging organisms in complex human microbiomes. Anaerobe 17, 56-63. Dridi, B., Henry, M., Richet, H., Raoult, D. & Drancourt, M. (2012a). Age-related prevalence of Methanomassiliicoccus luminyensis in the human gut microbiome. APMIS 120, 773-777. Dridi, B., Fardeau, M. L., Ollivier, B., Raoult, D. & Drancourt, M. (2012b). Methanomassiliicoccus luminyensis gen. nov., sp. nov., a methanogenic archaeon isolated from human faeces. Int J Syst Evol Microbiol 62, 1902-1907. Dridi, B., Khelaifia, S., Fardeau, M. L., Ollivier, B. & Drancourt, M. (2012c). Tungsten-enhanced growth of Methanosphaera stadtmanae. BMC Res Notes 5, 238. Fricke, W. F., Seedorf, H., Henne, A., Krüer, M., Liesegang, H., Hedderich, R., Gottschalk, G. & Thauer, R. K. (2006). The genome sequence of Methanosphaera stadtmanae reveals why this human intestinal archaeon is restricted to methanol and H2 for methane formation and ATP synthesis. J Bacteriol 188, 642-658. Gaci, N., Borrel, G., Tottey, W., O’Toole, P. W. & Brugère, J. F. (2014). Archaea and the human gut: new beginning of an old story. World J Gastroenterol 20, 16062-16078. Gibson, G. R., Macfarlane, G. T. & Cummings, J. H. (1993). Sulphate reducing bacteria and hydrogen metabolism in the human large intestine. Gut 34, 437-439. Hansen, E. E., Lozupone, C. A., Rey, F. E., Wu, M., Guruge, J. L., Narra, A., Goodfellow, J., Zaneveld, J. R., McDonald, D. T., Goodrich, J. A., Heath, A. C., Knight, R. & Gordon, J. I. (2011). Pan-genome of the dominant human gut-associated archaeon, Methanobrevibacter smithii, studied in twins. Proc Natl Acad Sci U S A 108 Suppl 1, 4599-4606. Jones, W. J., Nagle, D. P. Jr & Whitman, W. B. (1987). Methanogens and the diversity of archaebacteria. Microbiol Rev 51, 135-177. Jones, B. V., Begley, M., Hill, C., Gahan, C. G. & Marchesi, J. R. (2008). Functional and comparative metagenomic analysis of bile salt hydrolase activity in the human gut microbiome. Proc Natl Acad Sci U S A 105, 13580-13585. Khelaifia, S. & Raoult, D. (2016). Haloferax massiliensis sp. nov., the first human-associated halophilic archaea. New Microbes New Infect 12, 96-98. Lagier, J. C., Khelaifia, S., Alou, M. T., Ndongo, S., Dione, N., Hugon, P., Caputo, A., Cadoret, F., Traore, S. I., Seck, E. H., Dubourg, G., Durand, G., Mourembou, G., Guilhot, E., Togo, A., Bellali, S., Bachar, D., Cassir, N., Bittar, F., Delerce, J., Mailhe, M., Ricaboni, D., Bilen, M., Dangui Nieko, N. P., Dia Badiane, N. M., Valles, C., Mouelhi, D., Diop, K., Million, M., Musso, D., Abrahão, J., Azhar, E. I., Bibi, F., Yasir, M., Diallo, A., Sokhna, C., Djossou, F., Vitton, V., Robert, C., Rolain, J. M., La Scola, B., Fournier, P. E., Levasseur, A. & Raoult, D. (2016). Culture of previously uncultured members of the human gut microbiota by culturomics. Nat Microbiol 1, 16203. Levitt, M. D., Furne, J. K., Kuskowski, M. & Ruddy, J. (2006). Stability of human methanogenic flora over 35 years and a review of insights obtained from breath methane measurements. Clin Gastroenterol Hepatol 4, 123-129. Mihajlovski, A., Doré, J., Levenez, F., Alric, M. & Brugère, J. F. (2010). Molecular evaluation of the human gut methanogenic archaeal microbiota reveals an age-associated increase of the diversity. Environ Microbiol Rep 2, 272-280. Miller, T. L. & Wolin, M. J. (1982). Enumeration of Methanobrevibacter smithii in human feces. Arch Microbiol 131, 14-18. Oxley, A. P., Lanfranconi, M. P., Würdemann, D., Ott, S., Schreiber, S., McGenity, T. J., Timmis, K. N. & Nogales, B. (2010). Halophilic archaea in the human intestinal mucosa. Environ Microbiol 12, 2398-2410. Samuel, B. S., Hansen, E. E., Manchester, J. K., Coutinho, P. M., Henrissat, B., Fulton, R., Latreille, P., Kim, K., Wilson, R. K. & Gordon, J. I. (2007). Genomic and metabolic adaptations of Methanobrevibacter smithii to the human gut. Proc Natl Acad Sci U S A 104, 10643-10648. Söllinger, A., Schwab, C., Weinmaier, T., Loy, A., Tveit, A. T., Schleper, C. & Urich, T. (2016). Phylogenetic and genomic analysis of Methanomassiliicoccales in wetlands and animal intestinal tracts reveals clade-specific habitat preferences. FEMS Microbiol Ecol doi:10.1093/femsec/fiv149. Vanderhaeghen, S., Lacroix, C. & Schwab, C. (2015). Methanogen communities in stools of humans of different age and health status and co-occurrence with bacteria. FEMS Microbiol Lett 362, fnv092. Comments are closed.
|
Lesley HoylesProfessor of Microbiome and Systems Biology, Nottingham Trent University ArchivesCategories
All
|
